OAK Basics
OAK is a library for accessing and working with Ontologies, controlled vocabularies, and terminologies (from here on we will simply use “Ontology” in a very inclusive sense).
OAK is written in Python, and can be used in Python programs, or from the Command Line. Command line usage doesn’t require any programming experience or programming knowledge, but some knowledge of unix conventions might be helpful.
If you are looking for a quickstart guide to command line interfaces in general, see the Command line short intro.
What is an ontology and why would I want to access one?
Ontologies, controlled vocabularies, terminologies, and taxonomies are all ways of organizing data, information, and knowledge. Although OAK stands for Ontology Access Kit, we use the term “ontology” to refer to any of these types of knowledge organization.
An ontology typically consists of a set of Terms (also known as concepts, or classes) that serve as descriptors. For example, the Environment Ontology consists of classes representing different environments or environmental features, such as glaciers, lakes, and oceans.
These terms are often hierarchically organized, and may include other kinds of hierarchical information.
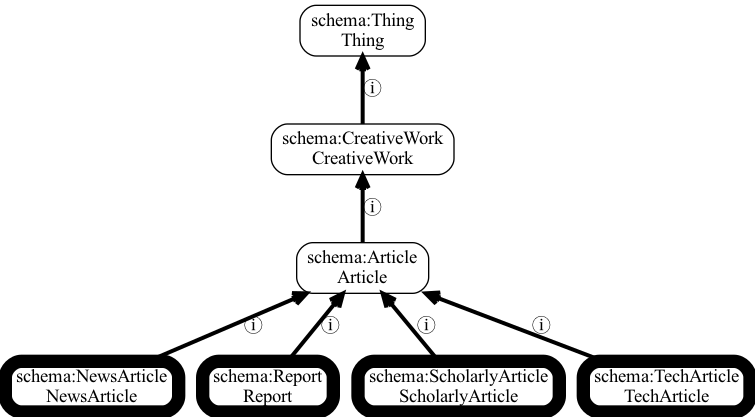
Example of a part of a vocabulary: Schema.org article types.
This example shows a small example of the portion of schema.org with some types of article. The vocabulary is organized in a hierarchy, with hierarchical relationships shown where ⓘ denotes an is-a Relationship. Note that although schema.org is more of a vocabulary than an ontology, you can use OAK to access it. This example was drawn using the OAK “viz” command.
Ontologies are used for many different tasks, but one of the most common ones is to annotate or “tag” pieces of data or data elements. For example, schema.org can be used to tag web pages, the Human Phenotype Ontology is used to annotate individual patients with a set of phenotypic features. In more formal ontology systems, the classes in an ontology are used to type instances or individuals. And in fact, an ontology can contain individuals too.
Ontologies may also include rich lexical information or other Metadata. These include textual Definitions, to help guide curators or users of the ontology, or aliases to help guide search.
Note
For more information on ontologies, see the Introduction to Ontologies in the OBO Academy Book.
Ontology languages, standards, and formats
The Web Ontology Language (OWL) is a standard for modeling ontologies. OWL is not a format in itself, but it has a number of different serializations, including RDF/XML, RDF/turtle, functional syntax, and Manchester Syntax.
It is not necessary to understand OWL to use OAK for many tasks. In fact, OWL is not the only way in which ontologies and vocabularies are represented.
Many biological ontologies have historically used OBO-Format, a simple text format. OBO-Format is mapped to and defined in terms of OWL.
A lot of terminologies and thesauri are represented in SKOS, a standard for representing controlled vocabularies.
Clinical vocabularies and ontologies are often represented as FHIR concepts or via formats like RRF
Some vocabularies such as schema.org use a lightweight RDF Schema representation.
OAK is able to read (and in some cases write) ontologies in all of these formats. Different Implementations handle different formats and backends. You can use Ontology Adapter Selectors to choose which implementation to use (alternatively, this is done automatically based on the file extension).
For example, on the command line, to list all terms a downloaded OBO Format file of the Human Phenotype Ontology:
$ runoak -i hp.obo terms
...
HP:0005671 ! Bilateral intracerebral calcifications
HP:0005676 ! Rudimentary postaxial polydactyly of hands
HP:0005678 ! Anterior atlanto-occipital dislocation
...
Note that each term has its own unique Identifier (in this case, a CURIE).
If you prefer to do this in Python rather than the command line:
>>> from oaklib import get_adapter
>>> hpo = get_adapter("hp.obo")
>>> for term_id in hpo.entities():
... print(term_id, hpo.label(term_id))
Even when ontologies are represented in OWL, it doesn’t mean that everything is standardized. Two OWL ontologies could choose to indicate primary Labels, synonyms, mappings, and definitions in different ways. OWL provides a framework for standardizing on the meaning of logical Axioms, but not on the different ways to represent metadata.
OAK attempts to provide a common layer on top of all these variants. See further sections of this guide for more details on how this is done.
Note
For more information on OWL and other ontology languages, see the section on OWL in the OBO Academy Book.
Core OAK concepts
Metadata
“other” logical constructs
In OAK, Entity is a general grouping for any identifiable thing. This includes Classes, Individuals, and Relationship Types, as well as ontologies themselves.